- Power BI forums
- Updates
- News & Announcements
- Get Help with Power BI
- Desktop
- Service
- Report Server
- Power Query
- Mobile Apps
- Developer
- DAX Commands and Tips
- Custom Visuals Development Discussion
- Health and Life Sciences
- Power BI Spanish forums
- Translated Spanish Desktop
- Power Platform Integration - Better Together!
- Power Platform Integrations (Read-only)
- Power Platform and Dynamics 365 Integrations (Read-only)
- Training and Consulting
- Instructor Led Training
- Dashboard in a Day for Women, by Women
- Galleries
- Community Connections & How-To Videos
- COVID-19 Data Stories Gallery
- Themes Gallery
- Data Stories Gallery
- R Script Showcase
- Webinars and Video Gallery
- Quick Measures Gallery
- 2021 MSBizAppsSummit Gallery
- 2020 MSBizAppsSummit Gallery
- 2019 MSBizAppsSummit Gallery
- Events
- Ideas
- Custom Visuals Ideas
- Issues
- Issues
- Events
- Upcoming Events
- Community Blog
- Power BI Community Blog
- Custom Visuals Community Blog
- Community Support
- Community Accounts & Registration
- Using the Community
- Community Feedback
Register now to learn Fabric in free live sessions led by the best Microsoft experts. From Apr 16 to May 9, in English and Spanish.
- Power BI forums
- Forums
- Get Help with Power BI
- Power Query
- Re: zoo's na.locf function not filling fields with...
- Subscribe to RSS Feed
- Mark Topic as New
- Mark Topic as Read
- Float this Topic for Current User
- Bookmark
- Subscribe
- Printer Friendly Page
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Permalink
- Report Inappropriate Content
zoo's na.locf function not filling fields with R script in Power Query Editor
I'm trying to use the zoo package's na.locf() function to fill missing values in adjacent rows. It works when I run it in RStudio, but it does not work in PowerBI. Here's an abridged version of the R code:
library(dplyr)
library(zoo)
library(lubridate)
fillGuids <- clean %>%
arrange(User, Serial, desc(ClientTime_clean), MetaGuid, desc(Action)) %>%
group_by(User,Serial,date(ClientTime_clean)) %>%
mutate(MetaGuid_filled = zoo::na.locf(MetaGuid, na.rm=FALSE)) %>%
ungroup() %>% #fill all missing MetaGuids that come after first log by User/Serial/date
This is the desired output for the MetaGuid_filled field, derived from the MetaGuid field. However, the output from PowerBI is just what's in MetaGuid. No errors are thrown, but this is not the expected behavior of na.locf. Not to mention the exact same code works when using my RStudio IDE.
| MetaGuid_filled | MetaGuid | User | Serial | Action | ClientTime_clean |
| A123456 | A123456 | A | 1 | ViewDocument | 10/22/2019 |
| A123456 | A123456 | A | 1 | ViewDocument | 10/22/2019 |
| A123456 | A123456 | A | 1 | ViewDocument | 10/22/2019 |
| A123456 | A123456 | A | 1 | ViewDocument | 10/22/2019 |
| A123456 | A | 1 | ExecuteSearch | 10/22/2019 | |
| B987654 | B987654 | A | 1 | ViewDocument | 10/22/2019 |
| B987654 | B987654 | A | 1 | ViewDocument | 10/22/2019 |
| B987654 | A | 1 | ExecuteSearch | 10/22/2019 |
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Permalink
- Report Inappropriate Content
any update?
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Permalink
- Report Inappropriate Content
Hi rlu,
I am not familiar on R script, I will discuss this with outhers and try to reproduce this in my environment. I will inform you as soon as I get it
In addition, if you want to fill it up, why not use M code to achieve this goal
You could replace empty cell by null, then click Fill->down
let
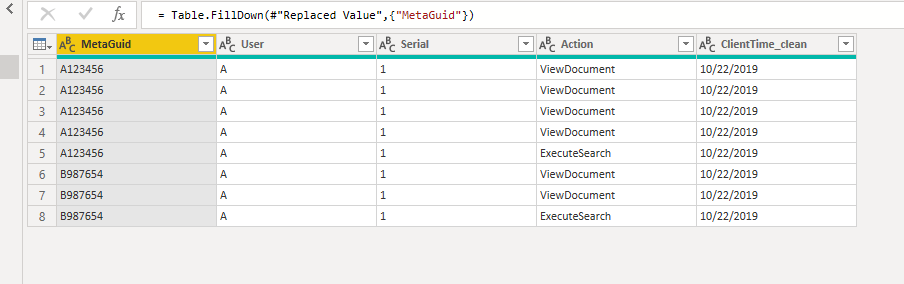
Source = Table.FromRows(Json.Document(Binary.Decompress(Binary.FromText("i45WcjQ0MjYxNVPSUXIEYkMgDstMLXfJTy7NTc0rAQkZ6BsZ6RsZGFoqxeoMPvVICl0rUpNLS1KDUxOLkjMwVTpZWpibmZoQbTKp6ol0SSwA", BinaryEncoding.Base64), Compression.Deflate)), let _t = ((type text) meta [Serialized.Text = true]) in type table [MetaGuid = _t, User = _t, Serial = _t, Action = _t, ClientTime_clean = _t]),
#"Replaced Value" = Table.ReplaceValue(Source,"",null,Replacer.ReplaceValue,{"MetaGuid"}),
#"Filled Down" = Table.FillDown(#"Replaced Value",{"MetaGuid"})
in
#"Filled Down"
Thanks for your understanding and support.
Best Regards,
Zoe Zhi
If this post helps, then please consider Accept it as the solution to help the other members find it more quickly.
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Permalink
- Report Inappropriate Content
Thanks for your response Zoe. I'm looking to maintain just one data munging pipeline as I use the same R script for offline analysis as well and wanted to use the same script to turn on a PowerBI dashboard to monitor metrics. Thus, M code will not be beneficial for me.
Helpful resources

Microsoft Fabric Learn Together
Covering the world! 9:00-10:30 AM Sydney, 4:00-5:30 PM CET (Paris/Berlin), 7:00-8:30 PM Mexico City

Power BI Monthly Update - April 2024
Check out the April 2024 Power BI update to learn about new features.

| User | Count |
|---|---|
| 101 | |
| 52 | |
| 21 | |
| 12 | |
| 11 |